Development of Microbial Bioremediation Products from Mixed Bacterial Consortia for Fat Degradation in Wastewater
Main Article Content
Abstract
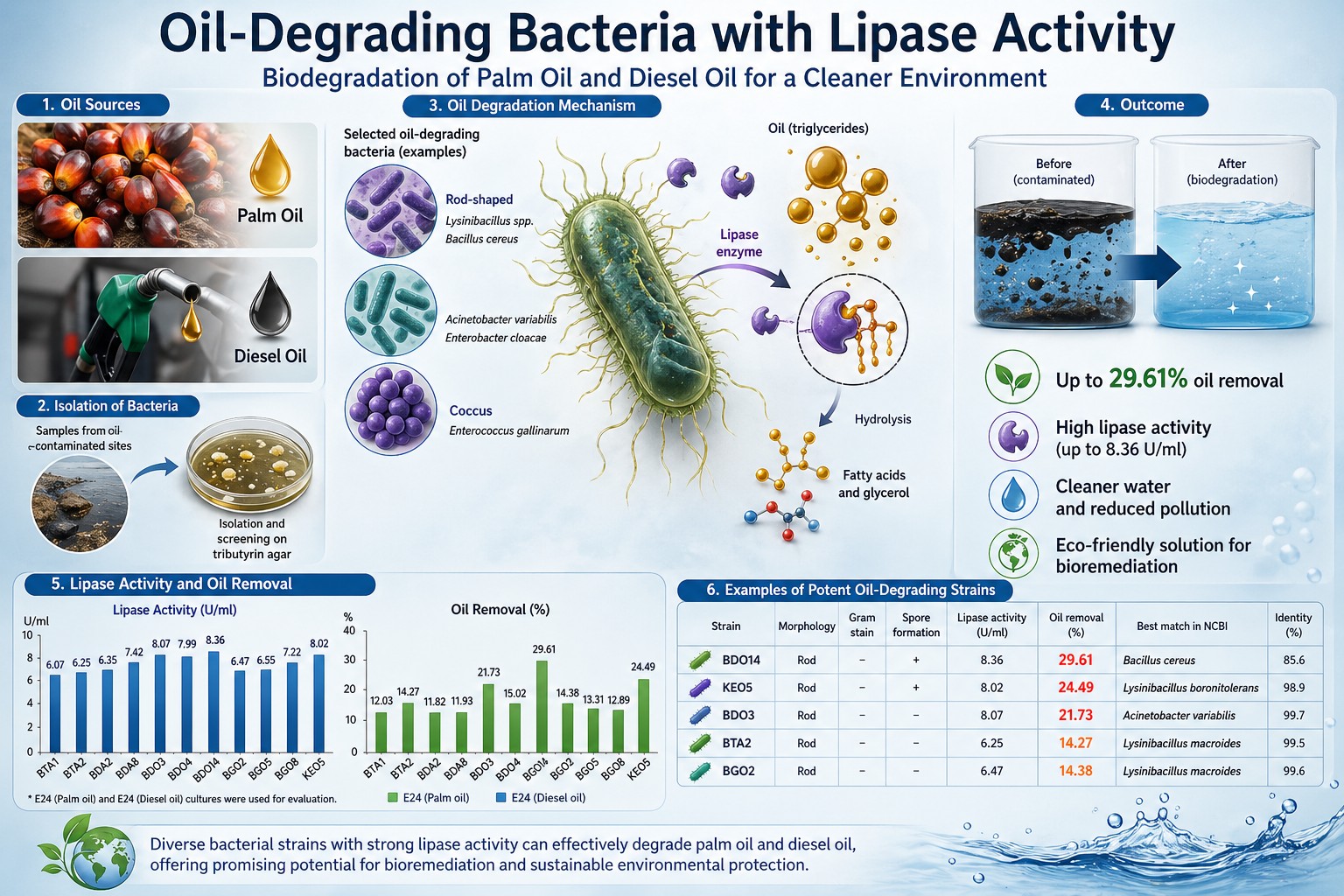
-rich wastewater from food-processing industries and municipalities causes pipeline blockage, odor, and impaired treatment performance. This study isolated, characterized, and formulated mixed bacterial consortia into bioremediation products and evaluated their efficacy at the laboratory and field scales. Eleven lipolytic isolates were recovered from high-fat wastewater sources by tributyrin agar screening; seven produced clear hydrolysis zones of 26–36 mm. Key strains identified by 16S rRNA gene sequencing were Acinetobacter variabilis BDO3 (lipase activity 8.07 U×mL⁻¹; oil removal 21.7%), Bacillus cereus BDO14 (8.36 U×mL⁻¹; 29.6%), and Lysinibacillus boronitolerans KEO5 (8.02 U×mL⁻¹; 24.5%). Nine consortium formulas were tested in simulated wastewater; the three-strain equal-ratio formula (BDO3:BDO14:KEO5 = 1:1:1) achieved the highest oil removal of 54.1% at day 6, representing an 83–149% improvement over the best monoculture. Scale-up fermentation to 200 L confirmed peak biomass at 48 h, with growth kinetics consistent with those at bench scale. Two product formats were developed: a rice bran–based powder retaining 2.45–6.0 × 10⁶ CFU×mL⁻¹ after 2 months, and a 15% glycerol liquid retaining 1.40–7.5 × 10⁷ CFU×mL⁻¹ after 1 month. Field applications of aquatic food-processing effluent, palm oil mill effluent, and municipal wastewater reduced residual fat to 28.7%, 48.8%, and 34.3%, respectively, within 6 days, outperforming two commercial products (residual fat 92–98%). The developed consortium-based bioremediation product demonstrates strong potential for sustainable, scalable management of fat wastewater at both community and industrial levels.
Article Details

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.
References
Wakelin, N. G.; Forster, C. F. An investigation into microbial removal of fats, oils and greases. Bioresour. Technol. 1997, 59, 37–43. https://doi.org/10.1016/S0960-8524(96)00134-4
Matos, A. T.; Lo Monaco, P. A.; Sarmento, A. P. Treatment of food-processing wastewater with high lipid content. Water Sci. Technol. 2018, 77, 1835–1843.
Marrakchi, M.; Hamza, W.; Sayadi, S. Electrocoagulation pre-treatment for lipid removal from olive mill wastewater. Sep. Purif. Technol. 2017, 182, 88–97.
Cammarota, M. C.; Freire, D. M. G. A review on hydrolytic enzymes in the treatment of wastewater with high oil and grease content. Bioresour. Technol. 2006, 97, 2195–2210. https://doi.org/10.1016/j.biortech.2006.02.030
Lesuisse, E.; Schanck, K.; Colson, C. Purification and preliminary characterization of the extracellular lipase of Bacillus subtilis 168. Eur. J. Biochem. 1993, 216, 155–160. https://doi.org/10.1111/j.1432-1033.1993.tb18127.x
Matsumiya, Y.; Wakita, D.; Kimura, A.; Sanpa, S.; Kubo, M. Isolation and characterization of a lipid-degrading bacterium and its application to lipid-containing wastewater treatment. J. Biosci. Bioeng. 2007, 103, 325–330. https://doi.org/10.1263/jbb.103.325
Prasad, M. P.; Manjunath, K. Comparative study on biodegradation of lipid-rich wastewater using lipase producing bacterial species. Indian J. Biotechnol. 2011, 10, 121–124.
Mongkolthanaruk, B.; Dharmsthiti, S. Biodegradation of lipid-rich wastewater by a mixed bacterial consortium. Int. Biodeterior. Biodegrad. 2002, 50, 101–105. https://doi.org/10.1016/S0964-8305(02)00057-4
Silva, C. F.; Azevedo, L. C. B.; Cardoso, D. E. S. Evaluation of microbial consortia for lipid degradation in wastewater treatment. J. Environ. Manage. 2015, 157, 1–8.
Morgan, C. A.; Herman, N.; White, P. A.; Vesey, G. Preservation of micro-organisms by drying. J. Microbiol. Methods 2006, 66, 183–193. https://doi.org/10.1016/j.mimet.2006.02.017
Fabian, C.; Ju, Y. H. A review on rice bran protein: Its properties and extraction methods. Crit. Rev. Food Sci. Nutr. 2011, 51, 816–827. https://doi.org/10.1080/10408398.2010.482678
Miyamoto-Shinohara, Y.; Imaizumi, T.; Sukenobe, J.; Murakami, Y.; Kawamura, S.; Komatsu, Y. Survival rate of microbes after freeze-drying and long-term storage. Cryobiology 2000, 41, 251–255. https://doi.org/10.1006/cryo.2000.2282
Lesuisse, E.; Schanck, K.; Colson, C. Purification and characterization of extracellular lipase on tributyrin agar. Eur. J. Biochem. 1993, 216, 155–160. https://doi.org/10.1111/j.1432-1033.1993.tb18127.x
Sharma, R.; Chisti, Y.; Banerjee, U. C. Production, purification, characterization, and applications of lipases. Biotechnol. Adv. 2001, 19, 627–662. https://doi.org/10.1016/S0734-9750(01)00086-6
Rahman, K. S. M.; Gakpe, E. Production, characterisation and applications of biosurfactants. Biotechnology 2008, 7, 360–370. https://doi.org/10.3923/biotech.2008.360.370
Altschul, S. F.; Gish, W.; Miller, W.; Myers, E. W.; Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Stanbury, P. F.; Whitaker, A.; Hall, S. J. Principles of Fermentation Technology, 3rd ed.; Elsevier: Oxford, U.K., 2017.
Vassilev, N.; Vassileva, M.; Azcon, R. Immobilised cell technology applied in solubilization of insoluble inorganic phosphate. Bioresour. Technol. 2001, 79, 263–271. https://doi.org/10.1016/S0960-8524(01)00017-7
APHA. Standard Methods for the Examination of Water and Wastewater, 23rd ed.; American Public Health Association: Washington, DC, 2017.
Jiang, Y.; Wen, J.; Bai, J.; Jia, X.; Hu, Z. Biodegradation of crude oil by a novel strain of Acinetobacter. Bioresour. Technol. 2012, 102, 7753–7758.
Turnbull, P. C. B.; Kramer, J. M. Non-gastrointestinal Bacillus cereus infections: An analysis of exotoxin production by strains isolated over a two-year period. J. Clin. Pathol. 1983, 36, 1091–1096. https://doi.org/10.1136/jcp.36.10.1091
Das, N.; Chandran, P. Microbial degradation of petroleum hydrocarbon contaminants: An overview. Biotechnol. Res. Int. 2011, 2011, 941810. https://doi.org/10.4061/2011/941810
Ahmed, I.; Yokota, A.; Yamazoe, A.; Fujiwara, T. Proposal of Lysinibacillus boronitolerans gen. nov., sp. nov. Int. J. Syst. Evol. Microbiol. 2007, 57, 1117–1125. https://doi.org/10.1099/ijs.0.63867-0
Madigan, M. T.; Bender, K. S.; Buckley, D. H.; Sattley, W. M.; Stahl, D. A. Brock Biology of Microorganisms, 15th ed.; Pearson: New York, NY, 2018.
Nicholson, W. L.; Munakata, N.; Horneck, G.; Melosh, H. J.; Setlow, P. Resistance of Bacillus endospores. Microbiol. Mol. Biol. Rev. 2000, 64, 548–572. https://doi.org/10.1128/MMBR.64.3.548-572.2000
Gottschalk, G. Bacterial Metabolism, 2nd ed.; Springer: New York, NY, 1986. https://doi.org/10.1007/978-1-4612-1072-6.
Shuler, M. L.; Kargi, F. Bioprocess Engineering: Basic Concepts, 3rd ed.; Prentice Hall: Upper Saddle River, NJ, 2017.
Ruiz, C.; Pastor, F. I. J.; Diaz, P. Isolation of lipid- and polysaccharide-degrading micro-organisms from subtropical forest soil, and analysis of lipolytic strain Bacillus sp. CR-179. Lett. Appl. Microbiol. 2005, 40, 218–227. https://doi.org/10.1111/j.1472-765X.2005.01660.x
Poh, P. E.; Chong, M. F. Development of anaerobic digestion methods for palm oil mill effluent treatment. Bioresour. Technol. 2009, 100, 1–9. https://doi.org/10.1016/j.biortech.2008.06.022