Prevalence of AmpC β-Lactamase and Regulatory Genes (LysR, QseB, and QseC) in Multidrug-Resistant Klebsiella pneumoniae Clinical Isolates from Al-Hillah, Iraq
Main Article Content
Abstract
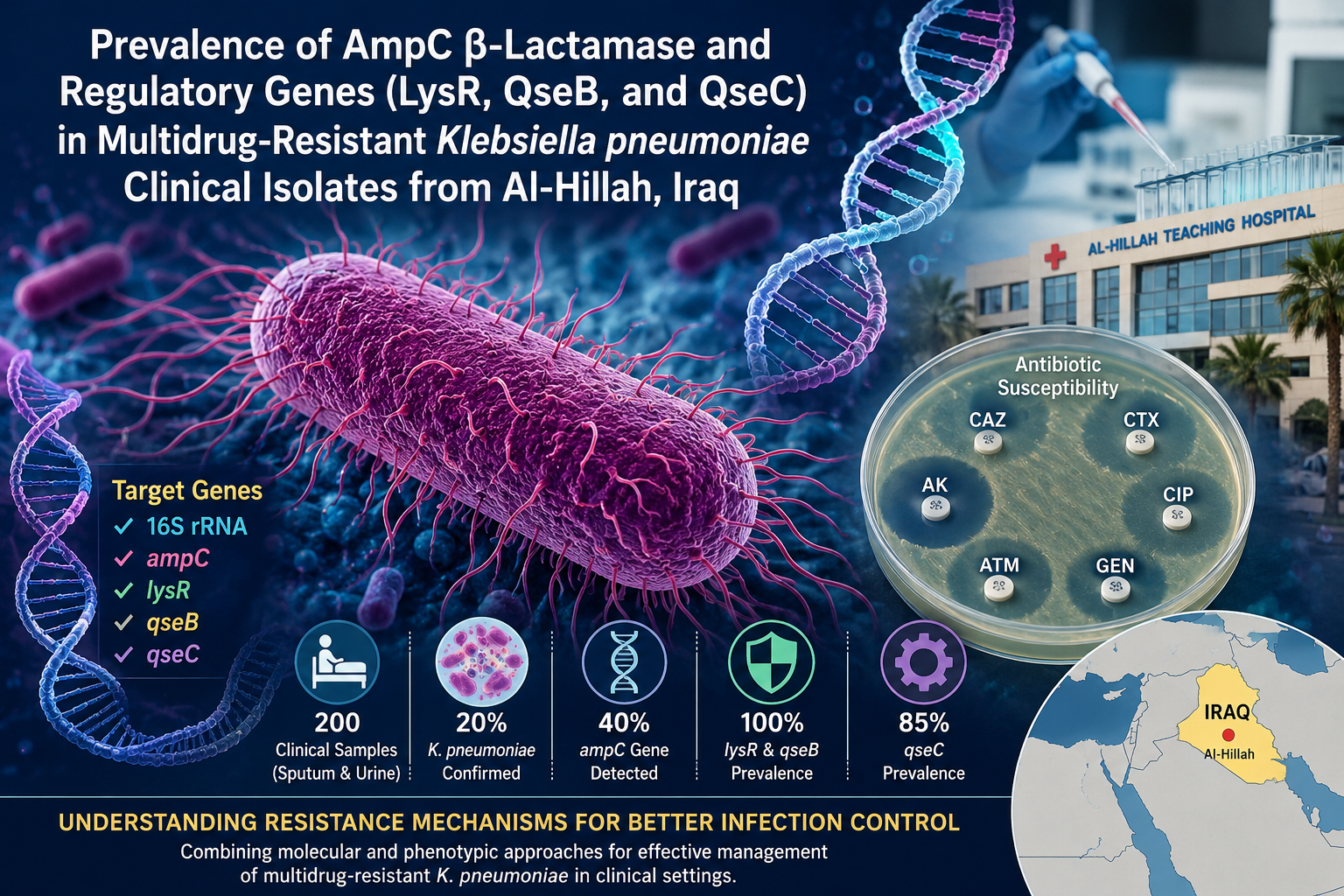
Klebsiella pneumoniae is a critical opportunistic pathogen classified by the World Health Organization among priority drug-resistant organisms, owing to its capacity to acquire multidrug resistance (MDR) and express diverse virulence mechanisms. Understanding the molecular basis of resistance and the prevalence of regulatory genes in clinical isolates is essential for effective infection control. A total of 200 clinical samples (sputum and urine) were collected from hospitalized patients at two teaching hospitals in Al-Hillah, Iraq. Isolates were identified using the VITEK 2 Compact system, and antibiotic susceptibility testing was performed. Genomic DNA was extracted by the boiling method, and conventional PCR was used to detect five target genes: 16S rRNA, ampC, lysR, qseB, and qseC. Of 100 culture-positive samples, twenty (20%) were confirmed as K. pneumoniae with 99% probability. High resistance rates were recorded against cephalosporins (100%) and ceftazidime/ceftriaxone (90%), while amikacin showed the highest susceptibility (85%). The ampC gene was detected in 40% of isolates. All isolates (100%) carried 16S rRNA, lysR, and qseB genes, while qseC was present in 85%. The widespread resistance—even in isolates lacking the ampC gene—suggests the possible involvement of additional concurrent mechanisms, such as ESBLs, efflux pumps, and porin loss, although these were not directly tested in the present study. The universal prevalence of regulatory genes lysR and qseB underscores their role in bacterial adaptation and virulence. Combined molecular and phenotypic approaches are essential for comprehensive characterization of MDR K. pneumoniae in clinical settings.
Article Details

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.
References
World Health Organization. Disease Outbreak News: Antimicrobial Resistance, Hypervirulent Klebsiella pneumoniae—Global Situation [Internet]. WHO: Geneva, 2024 Jul 31 [cited 2026 Mar 25]. Available from: https://www.who.int/emergencies/disease-outbreak-news/item/2024-DON527
Sukhumungoon, P.; Putchu, C.; Palee, T.; Bunkraiwong, P.; Yong-un, P.; Rattanachuay, P. Public Health Implications of Antimicrobial-Resistant Bacteria in the U-Tapao Canal, South of Thailand: A Study of Escherichia coli and Associated Gram-Negative Bacteria. ASEAN J. Sci. Technol. Rep. 2025, e260502. https://doi.org/10.55164/ajstr.v29i2.260502
Zubairi, R. B.; Abbas, B. A.; Abdullah, F. A. Surveillance of Rotavirus A Genotypes in Diarrheal Cases from Humans and Bovine in Basra Province, Iraq. ASEAN J. Sci. Technol. Rep. 2025, e261427.
Al-Jawahery, H. F.; Naji, N. S.; Al-Azawey, A. S. N. Physicochemical and Microbiological Evaluation of Reverse Osmosis Drinking Water Quality in Babylon Province, Iraq. ASEAN J. Sci. Technol. Rep. 2025, e260629.
Roberts, T.; Ling, C. L.; Watthanaworawit, W.; Cheav, C.; Sengduangphachanh, A.; Silisouk, J.; Hopkins, J.; Phommasone, K.; Batty, E. M.; Turner, P.; Ashley, E. A. AmpC β-Lactamases Detected in Southeast Asian Escherichia coli and Klebsiella pneumoniae. JAC-Antimicrob. Resist. 2024, 6, dlae195. https://doi.org/10.1093/jacamr/dlae195
Rao, A.; Naha, S.; Bhattacharjee, A.; Chattopadhyay, P.; Dutta, S.; Basu, S. Plasmid-Mediated AmpC in Klebsiella pneumoniae and Escherichia coli from Septicaemic Neonates: Diversity, Transmission and Phenotypic Detection. J. Glob. Antimicrob. Resist. 2023, 34, 9–14. https://doi.org/10.1016/j.jgar.2023.05.012
Raheem, R. R.; Jheel, W. A. W. Molecular Characterization and Integron Gene Prevalence in Bacterial Pathogens from Neonatal Omphalitis. ASEAN J. Sci. Technol. Rep. 2025, e261254.
Hennequin, C.; Forestier, C. oxyR, a LysR-Type Regulator Involved in Klebsiella pneumoniae Mucosal and Abiotic Colonization. Infect. Immun. 2009, 77, 5449–5457. https://doi.org/10.1128/IAI.00837-09
Zhang, R.; Zheng, Y.; Ding, C.; Wu, J.; Zhu, W.; Zhu, X.; Xu, G.; Chen, L. OxyR Contributes to the Oxidative Stress Capacity and Virulence of Hypervirulent Klebsiella pneumoniae ATCC 43816. Front. Cell. Infect. Microbiol. 2025, 15, 1661384. https://doi.org/10.3389/fcimb.2025.1661384
Al-Galebi, A. A. S.; Al-Hassani, M. K. A.; Kadhim, H. M.; Alkhudhairy, M. K. Phenotypic-Genotypic Characterization of Macrolide-Lincosamide-Streptogramin B Resistance in Staphylococcus saprophyticus UTI Isolates from Iraq. ASEAN J. Sci. Technol. Rep. 2025, e261225.
Fernandez-Ciruelos, B.; Potmis, T.; Solomin, V.; Wells, J. M. Cross-Talk between QseBC and PmrAB Two-Component Systems Is Crucial for Regulation of Motility and Colistin Resistance in Enteropathogenic Escherichia coli. PLoS Pathog. 2023, 19, e1011345. https://doi.org/10.1371/journal.ppat.1011345
Khajanchi, B. K.; Kozlova, E. V.; Sha, J.; Popov, V. L.; Chopra, A. K. The Two-Component QseBC Signalling System Regulates In Vitro and In Vivo Virulence of Aeromonas hydrophila. Microbiology 2012, 158, 259–271. https://doi.org/10.1099/mic.0.051805-0
Legese, M. H.; Asrat, D.; Aseffa, A.; Hasan, B.; Mihret, A.; Swedberg, G. Molecular Epidemiology of Extended-Spectrum Beta-Lactamase and AmpC Producing Enterobacteriaceae among Sepsis Patients in Ethiopia: A Prospective Multicenter Study. Antibiotics 2022, 11, 131. https://doi.org/10.3390/antibiotics11020131
Fu, S.; Wen, C.; Wang, Z.; Qiu, Y.; Zhang, Y.; Zuo, J.; Xu, Y.; Han, X.; Luo, Z.; Chen, W.; Miao, J. Molecular Epidemiology and Antimicrobial Resistance of Outbreaks of Klebsiella pneumoniae Clinical Mastitis in Chinese Dairy Farms. Microbiol. Spectr. 2022, 10, e02997-22. https://doi.org/10.1128/spectrum.02997-22
Liu, W.; Bao, Q.; Qing, M.; Chen, X.; Sun, T.; Li, M.; Zhang, J.; Yu, J.; Bilige, M.; Sun, T.; Zhang, H. Isolation and Identification of Lactic Acid Bacteria from Tarag in Eastern Inner Mongolia of China by 16S rRNA Sequences and DGGE Analysis. Microbiol. Res. 2012, 167, 110–115. https://doi.org/10.1016/j.micres.2011.05.001
Mohamed, I. Q.; Al-Taai, H. R. Phylogenetic Analysis of Klebsiella pneumoniae Isolated from Nosocomial and Community Infection in Diyala, Iraq. Iraqi J. Sci. 2023, 64, 2726–2740. https://doi.org/10.24996/ijs.2023.64.6.7
Hussein, A. M.; Jameel, M. I.; Majeed, N. R.; Noori, A. A.; Mohammed, M. M.; Hamad, W. N.; Mohammedsaeed, A. A. Correlation between Klebsiella Species Biofilm Formation and Antibiotic Resistance from Erbil Hospital Patients. Cihan Univ.-Erbil Sci. J. 2024, 8, 56–64. https://doi.org/10.24086/cuesj.v8n2y2024.pp56-64
Indoung, S.; Watcharakul, S. Bactericidal Properties, Biofilm Formation Inhibition, and Chemical Profiling of Piper argyrites and Piper betel L. ASEAN J. Sci. Technol. Rep. 2025, e259978. https://doi.org/10.55164/ajstr.v28i5.259978
Abdul Muhsin, Z. A.; Jihad, G. H.; Yaaqoob, L. A. Green Synthesis of ZnO-TiO2 Nanoparticles Using Allium ampeloprasum (Kurrat) Extract and Their Antibacterial Activity. ASEAN J. Sci. Technol. Rep. 2024, e256572. https://doi.org/10.55164/ajstr.v28i4.256572
Joda, H. M.; Hamza, A. K. Effects of Dietary Insect Powder Supplementation on Hematological Parameters of Common Carp (Cyprinus carpio) Fry. ASEAN J. Sci. Technol. Rep. 2025, e260795.
Altaweel, D. Enhancement of Bioactive Compounds and Nutrient Content in Rosemary (Salvia rosmarinus) Using Nano-Magnesium and NPK Fertilization: A GC-MS Analysis. ASEAN J. Sci. Technol. Rep. 2025, e261168.
Lertworapreecha, M. Cloning and Nucleotide Sequence Analysis of Complementary DNA Encoding Antimicrobial Peptide from Dung Beetle Larva. ASEAN J. Sci. Technol. Rep. 2022, 134–142.
Zahwar, A. T.; Sabour, A. N. Berberine Ameliorates Methionine-Induced Hyperhomocysteinemia and Biochemical Alterations in Male Rats. ASEAN J. Sci. Technol. Rep. 2025, e262094.
Mutar, B. A.; Sekhi, A. A.; Alfarhani, Z. A. O.; Muttar, A. A. Comparison of Inhibitory Effects between Cetuximab and Cisplatin on Colon Cancer SW480 Cell Line. ASEAN J. Sci. Technol. Rep. 2025, e262531.
Abdulhasan, G. A.; Fadhil, H. Y.; Jasem, K. A. Detection of Genes Encoding Extended-Spectrum and AmpC β-Lactamases in Klebsiella pneumoniae Isolates from Clinical Specimens. Al-Nahrain J. Sci. 2015, 18, 125–132. https://doi.org/10.22401/JNUS.18.2.16
Albassam, W. W. Detection of Antibiotics Resistance Genes in Clinical Isolates of Klebsiella pneumoniae. Iraqi J. Sci. 2015, 56, 407–416.
Philippon, A.; Arlet, G.; Labia, R.; Iorga, B. I. Class C β-Lactamases: Molecular Characteristics. Clin. Microbiol. Rev. 2022, 35, e00150-21. https://doi.org/10.1128/cmr.00150-21
Badamchi, A.; Shabestari, A. N.; Etemadi, M. R.; Tabatabaei, A. Molecular Epidemiology of blaCMY-1, blaCMY-2, blaFOX Genes in K. pneumoniae from Elderly Patients in Tehran, Iran. Acta Med. Iran. 2021, 59, 472–476. https://doi.org/10.18502/acta.v59i8.7250
Realegeno, S.; Ward, K.; Garner, O. B.; Yang, S. Deceiving Phenotypic Susceptibility Results on a Klebsiella pneumoniae Blood Isolate Carrying Plasmid-Mediated AmpC Gene blaDHA-1. Front. Cell. Infect. Microbiol. 2021, 11, 561880. https://doi.org/10.3389/fcimb.2021.561880
Monawer, A. T. Molecular Analysis of Klebsiella pneumoniae Isolates Collected from Sputum Samples in Duhok Iraq. Al-Nahrain J. Sci. 2025, 28. https://doi.org/10.22401/ANJS.28.1.10
Kumar, S.; Anwer, R.; Azzi, A. Molecular Typing Methods and Resistance Mechanisms of MDR Klebsiella pneumoniae. AIMS Microbiol. 2023, 9, 112. https://doi.org/10.3934/microbiol.2023008
Jomehzadeh, N.; Ahmadi, K.; Shaabaninejad, H.; Eslami, G. Plasmid-Mediated AmpC β-Lactamase Gene Analysis in Klebsiella pneumoniae Clinical Isolates. Biomed. Biotechnol. Res. J. 2022, 6, 582–585. https://doi.org/10.4103/bbrj.bbrj_302_22
Etemadi, S.; Leylabadlo, H. E.; Ghotaslou, R. AmpC β-Lactamase among Enterobacteriaceae: A New Insight. Gene Rep. 2020, 19, 100673. https://doi.org/10.1016/j.genrep.2020.100673
Srinivasan, V. B.; Mondal, A.; Venkataramaiah, M.; Chauhan, N. K.; Rajamohan, G. Role of oxyRKP, a Novel LysR-Family Transcriptional Regulator, in Antimicrobial Resistance and Virulence in Klebsiella pneumoniae. Microbiology 2013, 159, 1301–1314. https://doi.org/10.1099/mic.0.065052-0
Yang, W.; Wang, W. Y.; Zhao, W.; Cheng, J. G.; Wang, Y.; Yao, X. P.; Yang, Z. X.; Yu, D.; Luo, Y. Preliminary Study on the Role of Novel LysR Family Gene kp05372 in Klebsiella pneumoniae of Forest Musk Deer. J. Zhejiang Univ.-Sci. B 2020, 21, 137–154. https://doi.org/10.1631/jzus.B1900440
Baugh, A. C.; Momany, C.; Neidle, E. L. Versatility and Complexity: Common and Uncommon Facets of LysR-Type Transcriptional Regulators. Annu. Rev. Microbiol. 2023, 77, 317–339. https://doi.org/10.1146/annurev-micro-050323-040543
Zhu, Y.; Dou, Q.; Du, L.; Wang, Y. QseB/QseC: A Two-Component System Globally Regulating Bacterial Behaviors. Trends Microbiol. 2023, 31, 749–762. https://doi.org/10.1016/j.tim.2023.02.001
Lv, J.; Zhu, J.; Wang, T.; Xie, X.; Wang, T.; Zhu, Z.; Chen, L.; Zhong, F.; Du, H. The Role of the Two-Component QseBC Signaling System in Biofilm Formation and Virulence of Hypervirulent Klebsiella pneumoniae ATCC43816. Front. Microbiol. 2022, 13, 817494. https://doi.org/10.3389/fmicb.2022.817494
Chu, X.; Yang, Q. Regulatory Mechanisms and Physiological Impacts of Quorum Sensing in Gram-Negative Bacteria. Infect. Drug Resist. 2024, 5395–5410. https://doi.org/10.2147/IDR.S485388
Sharma, S.; Kumar, S.; Kumar, P.; Tripathi, V. N. Quorum Sensing in Gram-Negative Pathogens, a Fresh Look. Microbe 2024, 4, 100108. https://doi.org/10.1016/j.microb.2024.100108
Zorgani, A.; Daw, H.; Sufya, N.; Bashein, A.; Elahmer, O.; Chouchani, C. Co-Occurrence of Plasmid-Mediated AmpC β-Lactamase Activity among Klebsiella pneumoniae and Escherichia coli. Open Microbiol. J. 2017, 11, 195–202. https://doi.org/10.2174/1874285801711010195
Faghihi, K.; Tajbakhsh, S.; Fouladvand, M.; Latifi, B.; Yousefi, F. Detection of Plasmid-Mediated AmpC β-Lactamases in Klebsiella pneumoniae Clinical Isolates from Bushehr Province, Iran. Iran. J. Microbiol. 2023, 15, 373–380. https://doi.org/10.18502/ijm.v15i3.12897
Barceló, I. M.; Escobar-Salom, M.; Cabot, G.; Perelló-Bauzà, P.; Jordana-Lluch, E.; Taltavull, B.; Torrens, G.; Rojo-Molinero, E.; Zamorano, L.; Pérez, A.; Oliver, A. Transferable AmpCs in Klebsiella pneumoniae: Interplay with Peptidoglycan Recycling, Mechanisms of Hyperproduction, and Virulence Implications. Antimicrob. Agents Chemother. 2024, 68, e01315-23. https://doi.org/10.1128/aac.01315-23
Li, G.; Wang, L.; Zhang, H.; Luan, Y.; Sun, Q.; Duo, L. Study on the Role of ampG in the Regulation of Plasmid-Mediated ampC-Induced Expression in Klebsiella pneumoniae. Infect. Drug Resist. 2023, 5587–5598. https://doi.org/10.2147/IDR.S421598